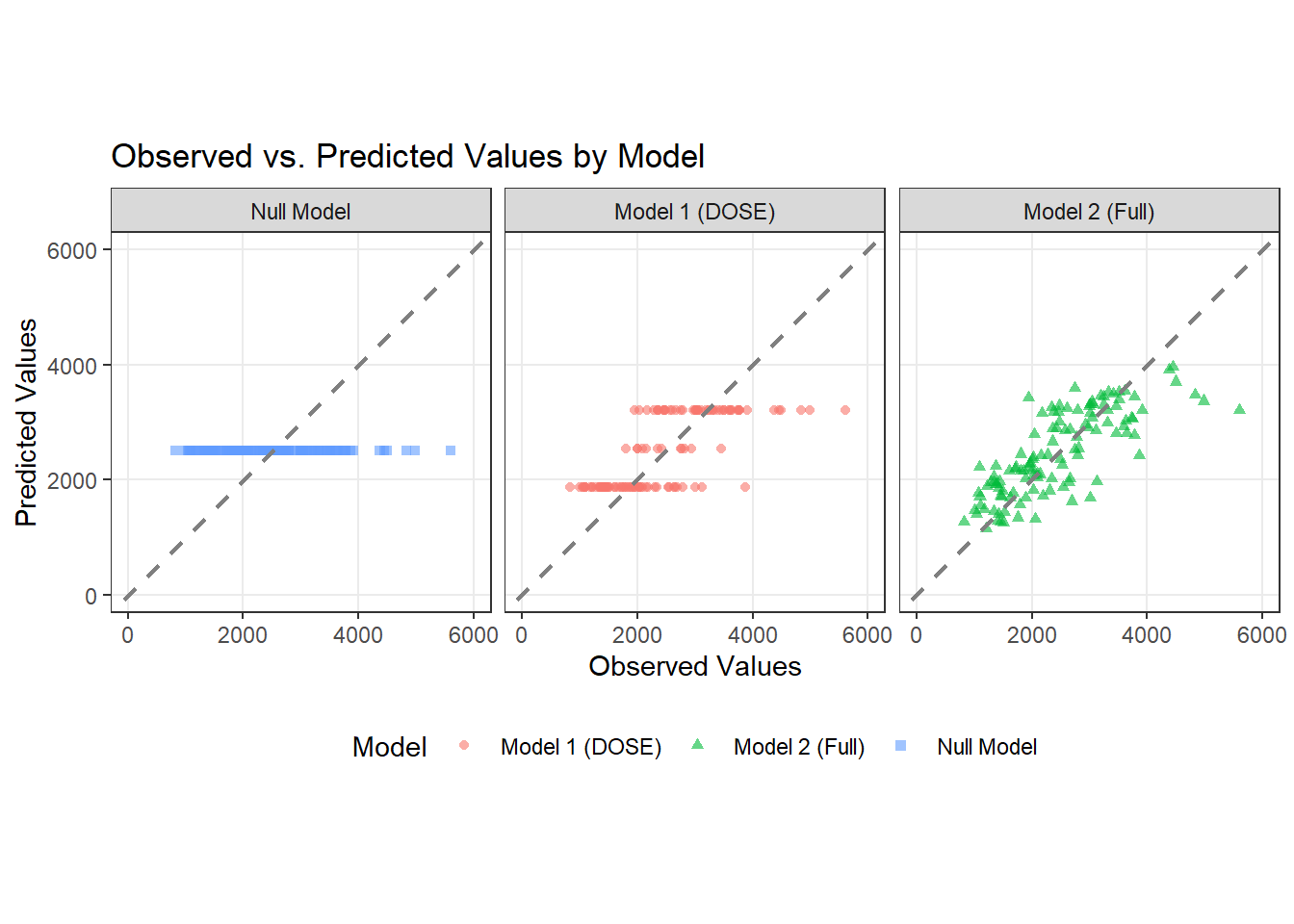
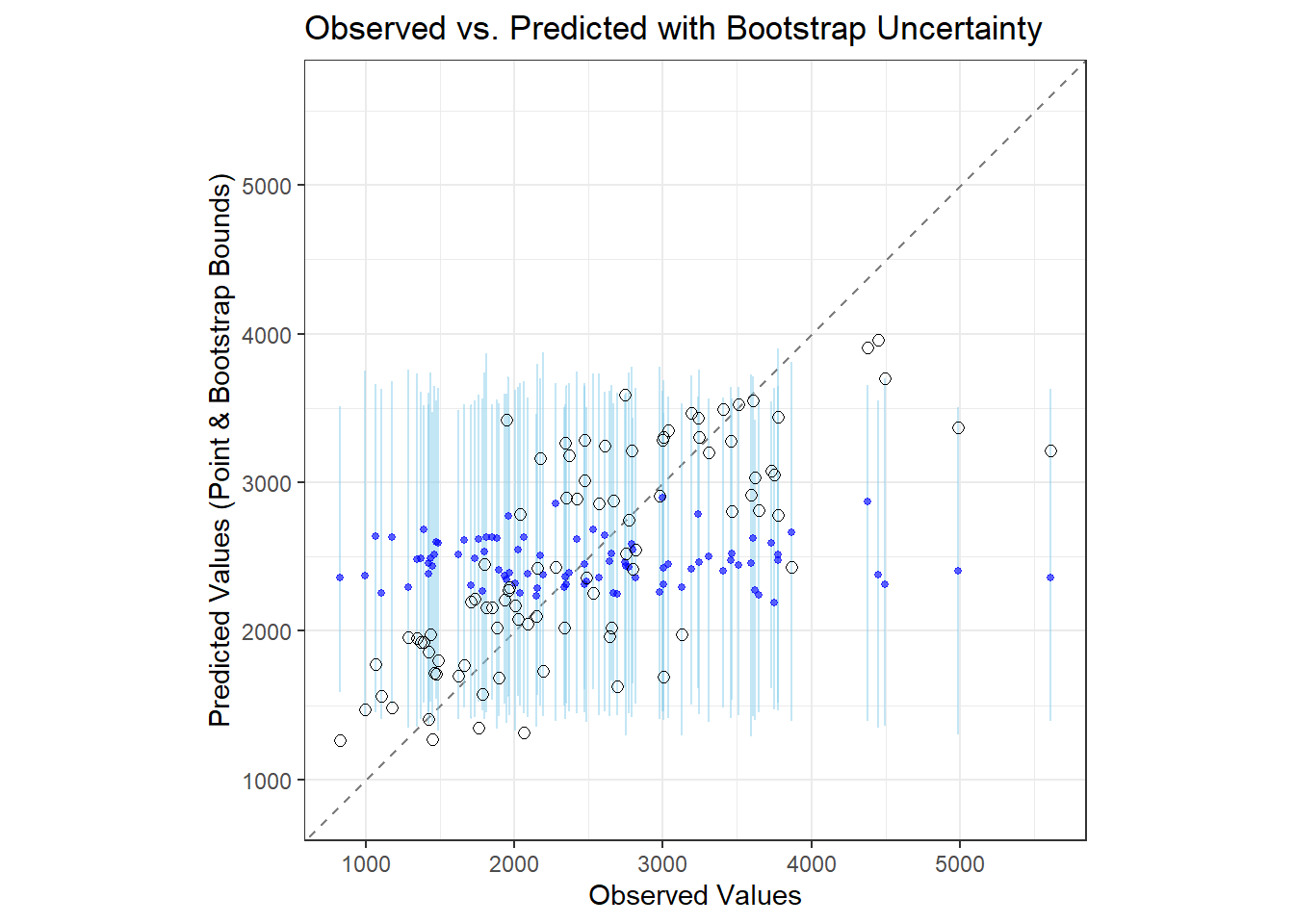
# Plotting observed vs predicted
ggplot(pred_long_format, aes(x = Y, y = prediction, color = model, shape = model)) +
geom_point(alpha = 0.6, size = 1.5) +
geom_abline(slope = 1, intercept = 0, color = "gray50", linewidth = 0.8, linetype = "dashed") +
scale_x_continuous(limits = c(0, 6000)) + # englarged to 6000 because there were values outside the plot
scale_y_continuous(limits = c(0, 6000)) + # englarged to 6000 because there were values outside the plot
coord_fixed(ratio = 1) + # Ensures 45° line appears at true 45°
labs(
title = "Observed vs. Predicted Values by Model",
x = "Observed Values",
y = "Predicted Values",
color = "Model",
shape = "Model"
) +
theme_bw() +
theme(
panel.grid.minor = element_blank(),
legend.position = "bottom"
) +
facet_wrap(~ factor(model,
levels = c("Null Model", "Model 1 (DOSE)", "Model 2 (Full)")))