library(gt)
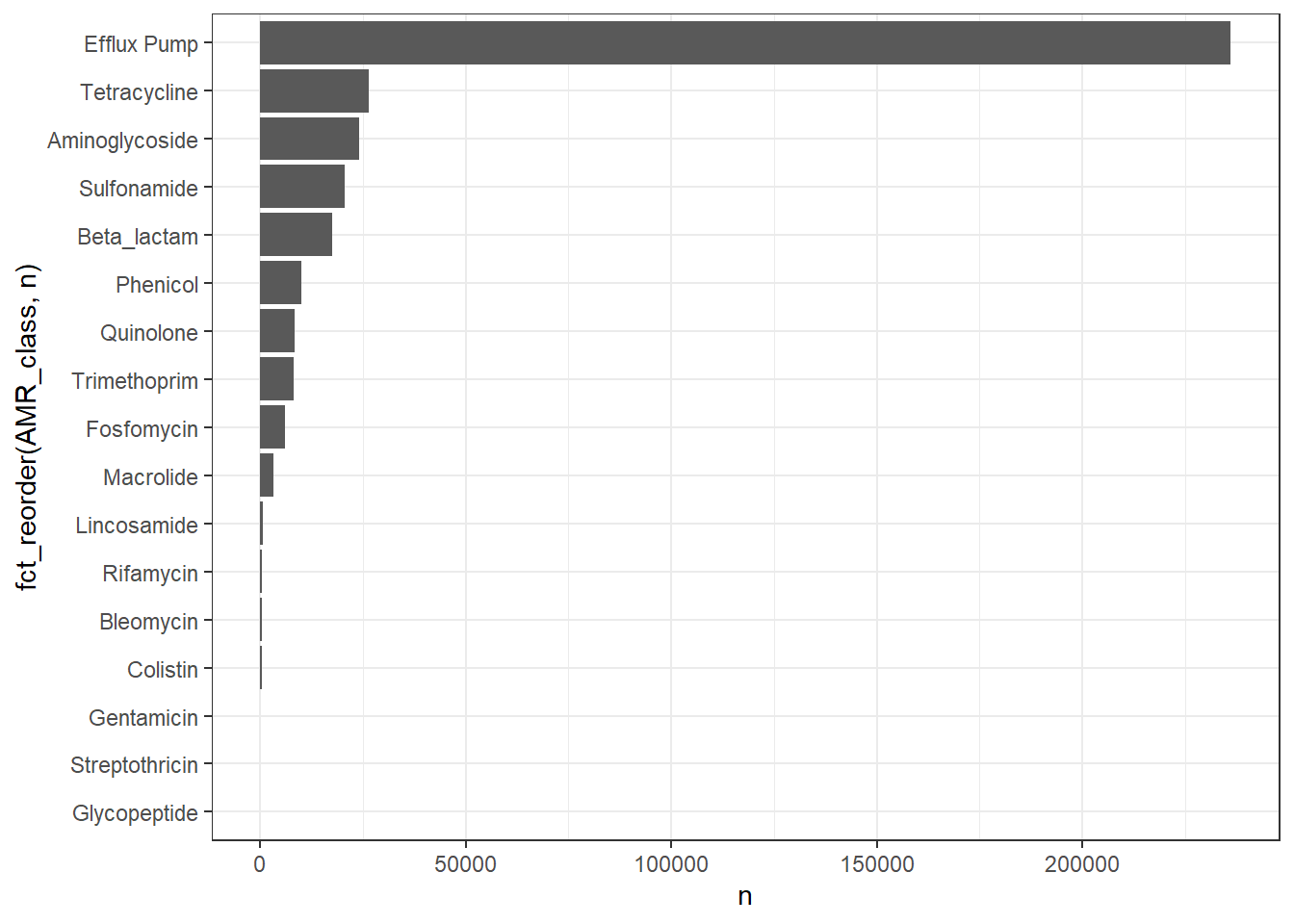
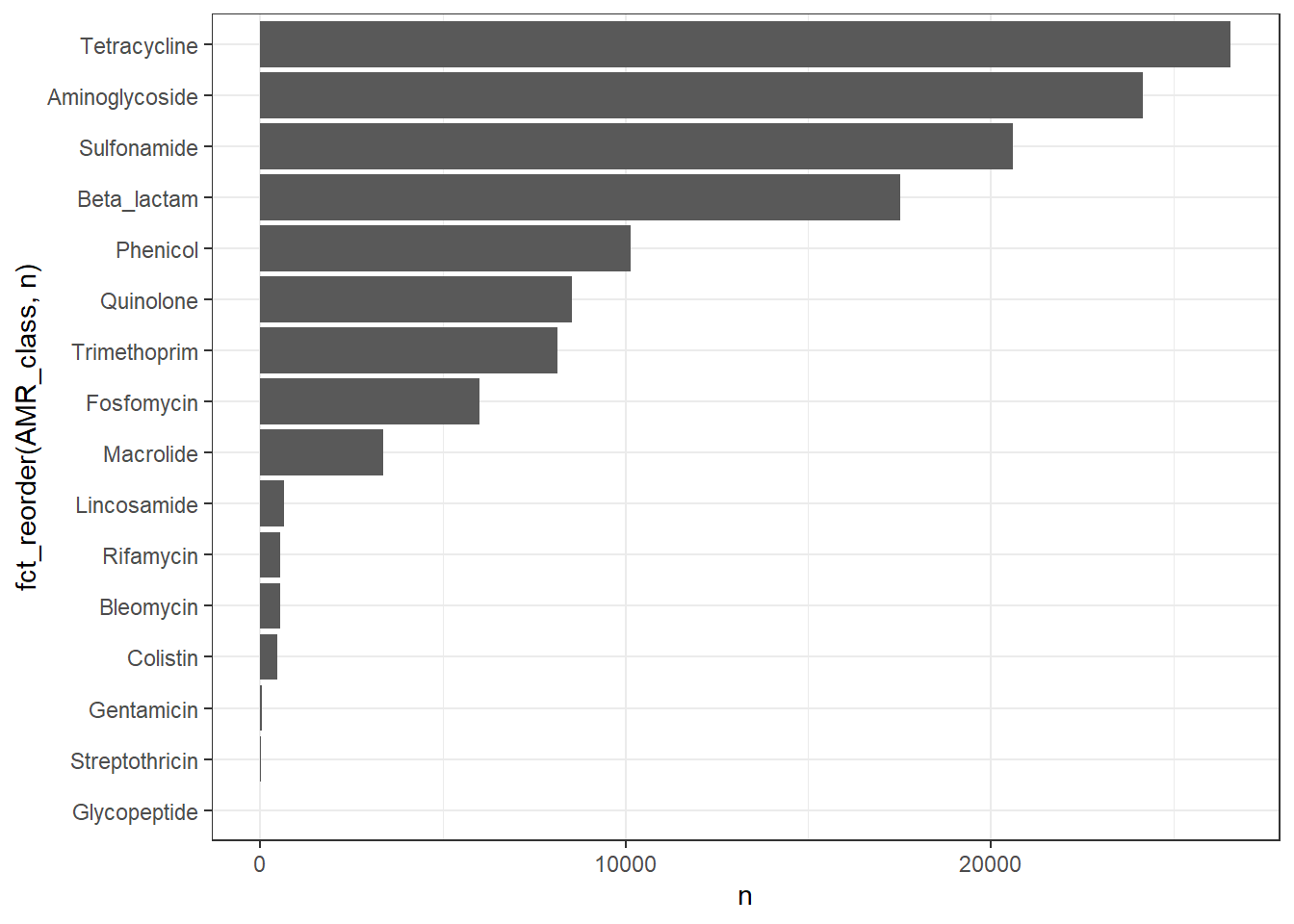
# Publication-ready Table for Class Prevalence
class_prev1 %>%
gt() %>%
tab_header(
title = "Prevalence of Clinically Relevant AMR Classes",
subtitle = "Excluding Efflux Pumps; (n = 245,772 isolates)"
) %>%
cols_label(
AMR_class = "Antimicrobial Class",
n = "Isolate Count",
Percentage = "Prevalence (%)"
) %>%
fmt_number(columns = n, decimals = 0, use_seps = TRUE) %>%
data_color(
columns = Percentage,
palette = "YlOrRd",
domain = c(0, 12)
) %>%
opt_stylize(style = 1, color = "gray") %>%
gtsave(here("results/tables/amr_class_prevalence.html"))
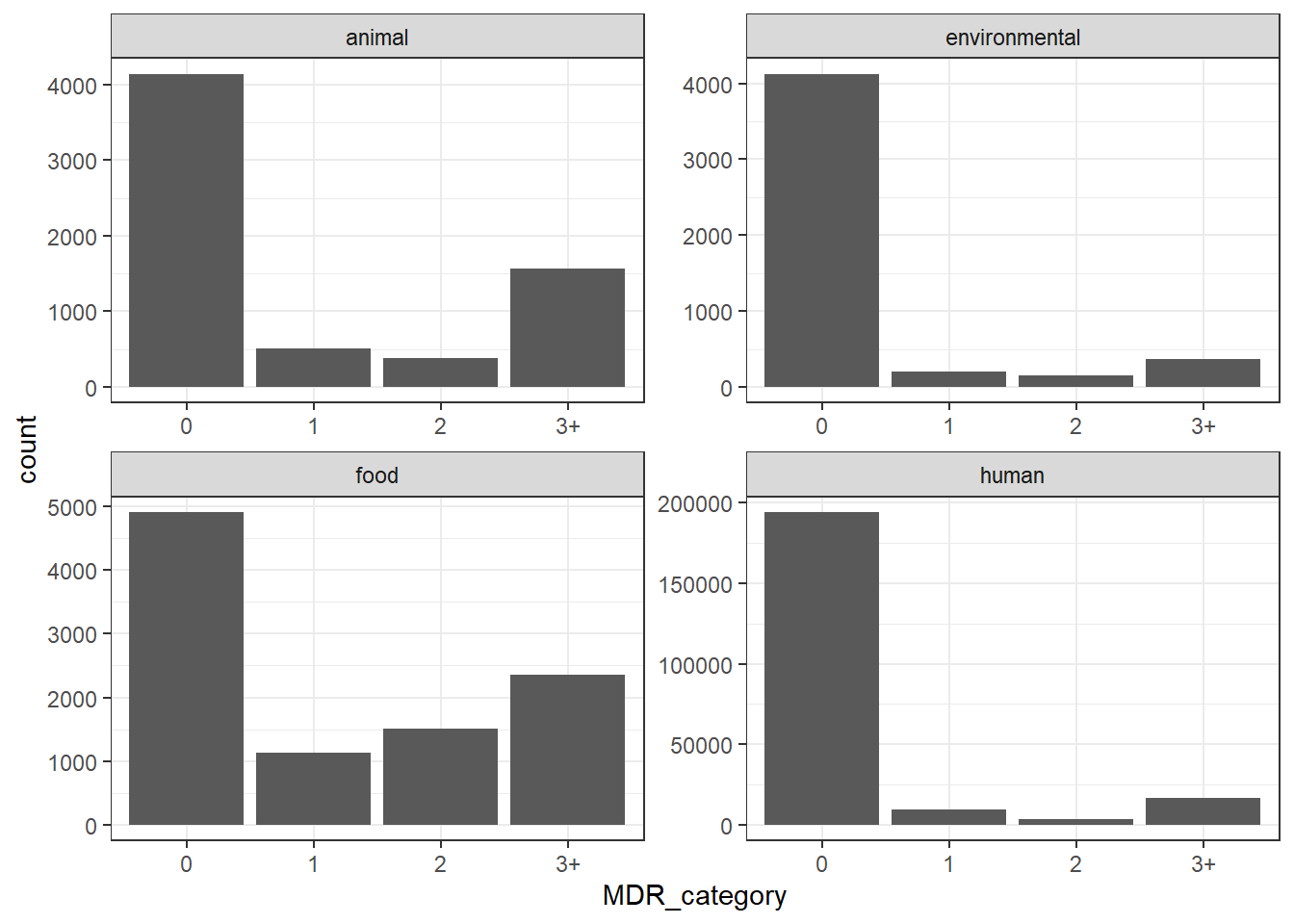
# Publication-ready MDR Plot
ggplot(isolates1, aes(x = MDR_category, fill = MDR_category)) +
geom_bar(show.legend = FALSE) +
facet_wrap(~ Source.type, scales = "free_y") +
scale_fill_brewer(palette = "Blues") +
labs(
title = "Distribution of Multidrug Resistance (MDR) by Source",
subtitle = "Categories: 0, 1, 2, or 3+ resistance classes",
x = "Number of Resistance Classes",
y = "Isolate Count"
) +
theme_bw() +
theme(strip.background = element_rect(fill = "white"),
strip.text = element_text(face = "bold")) -> amr_graph2_pub
ggsave(here("results/figures/amr_mdr_distribution.png"), plot = amr_graph2_pub, dpi = 300, width = 8, height = 6)
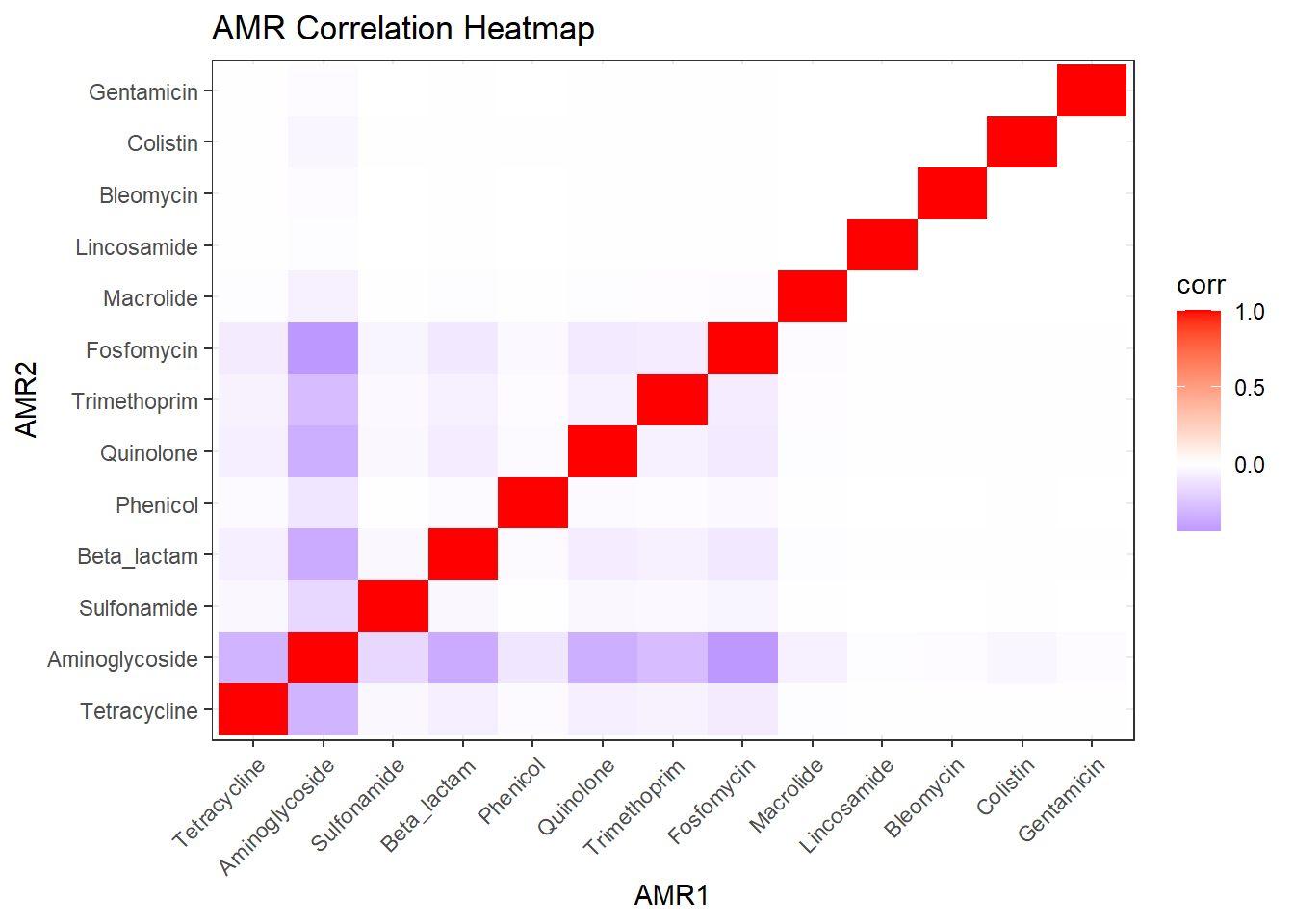
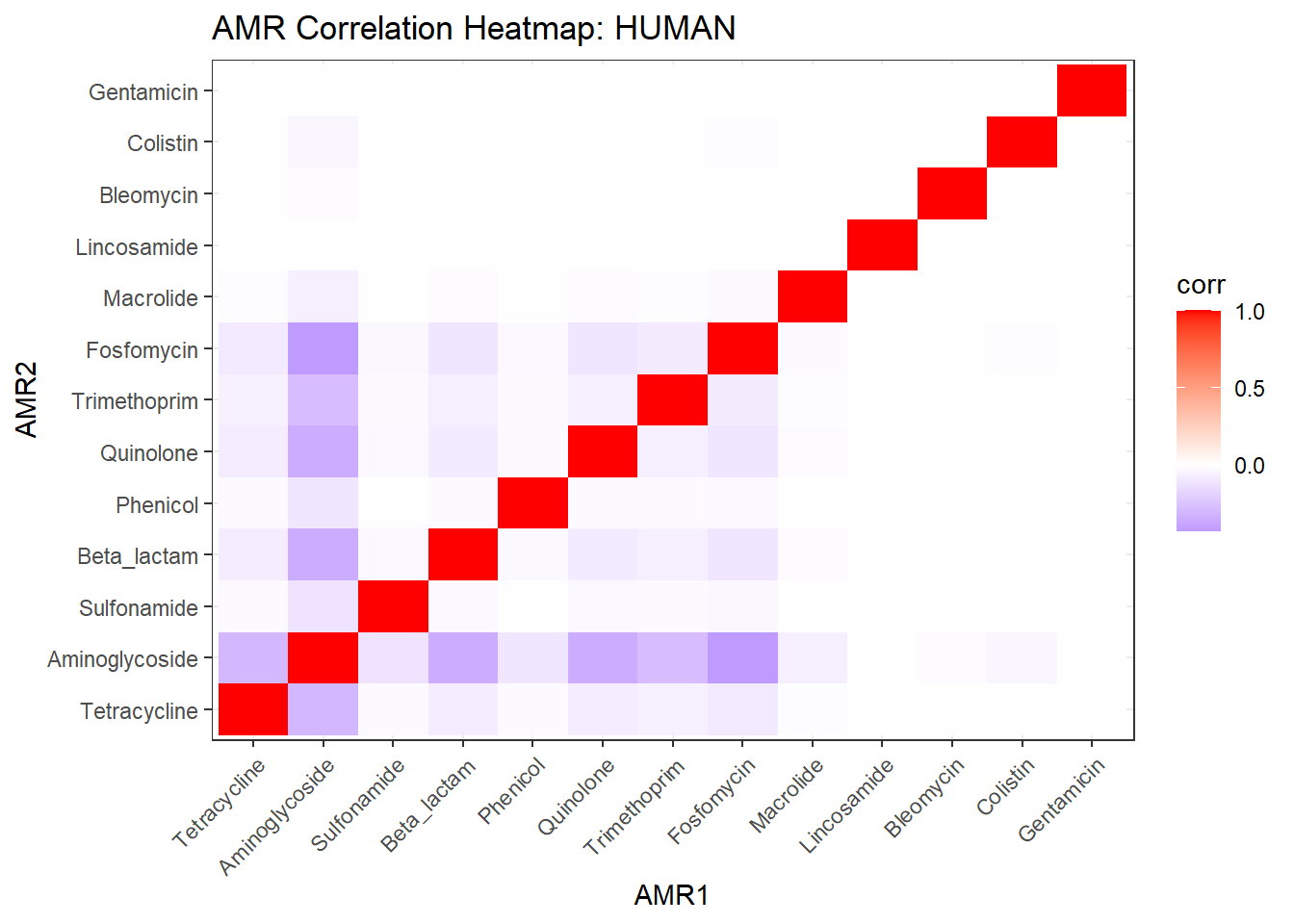
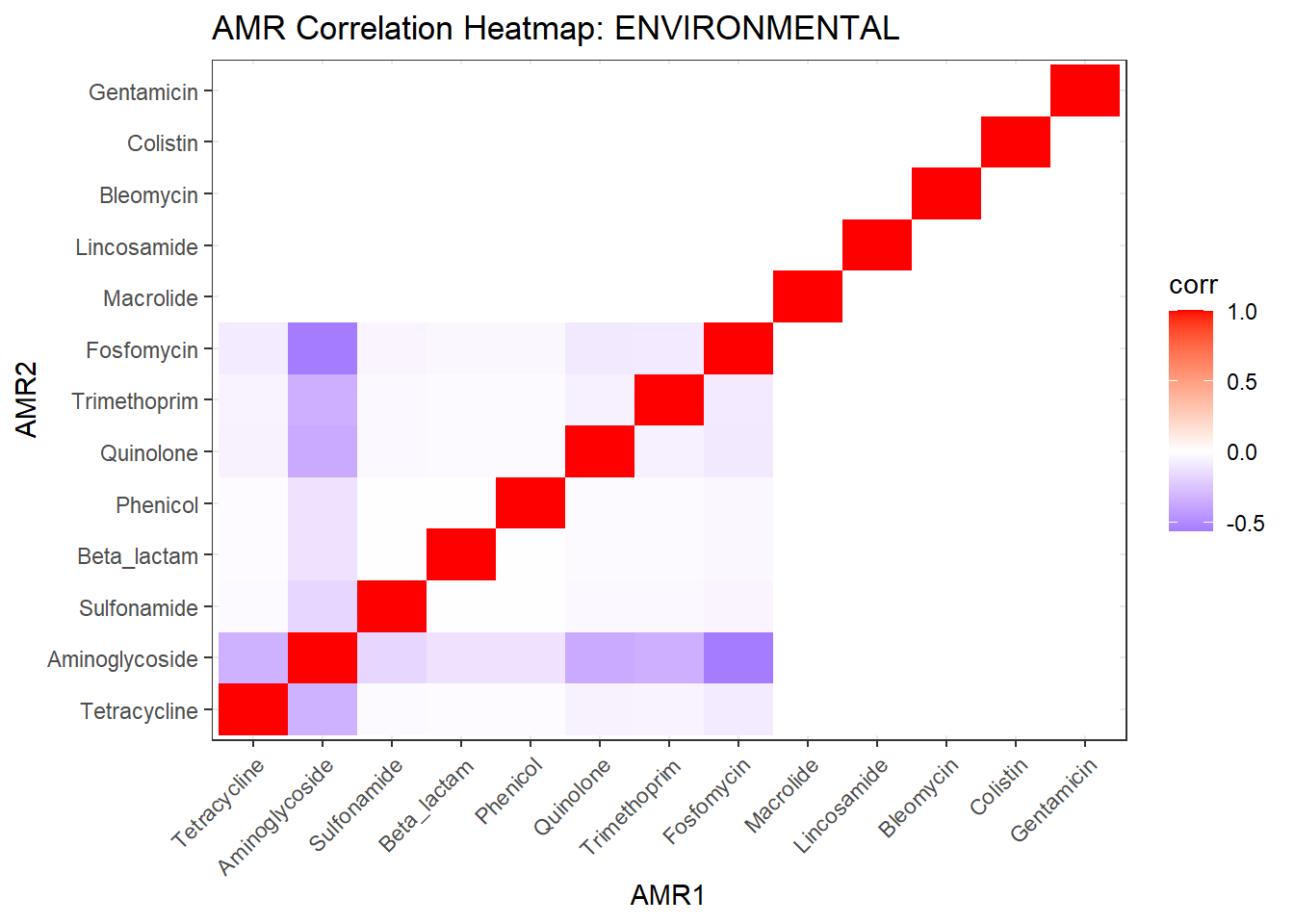
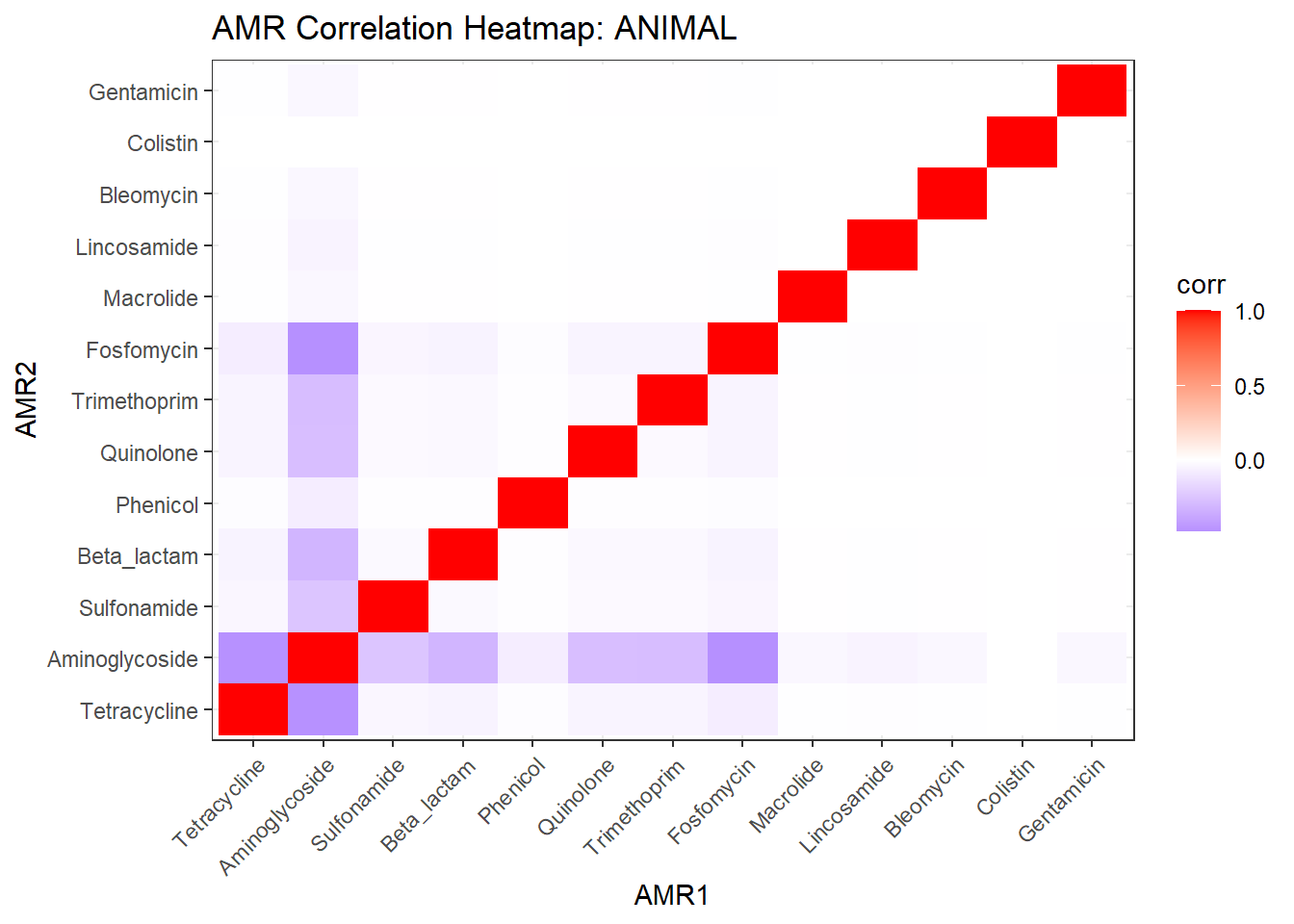
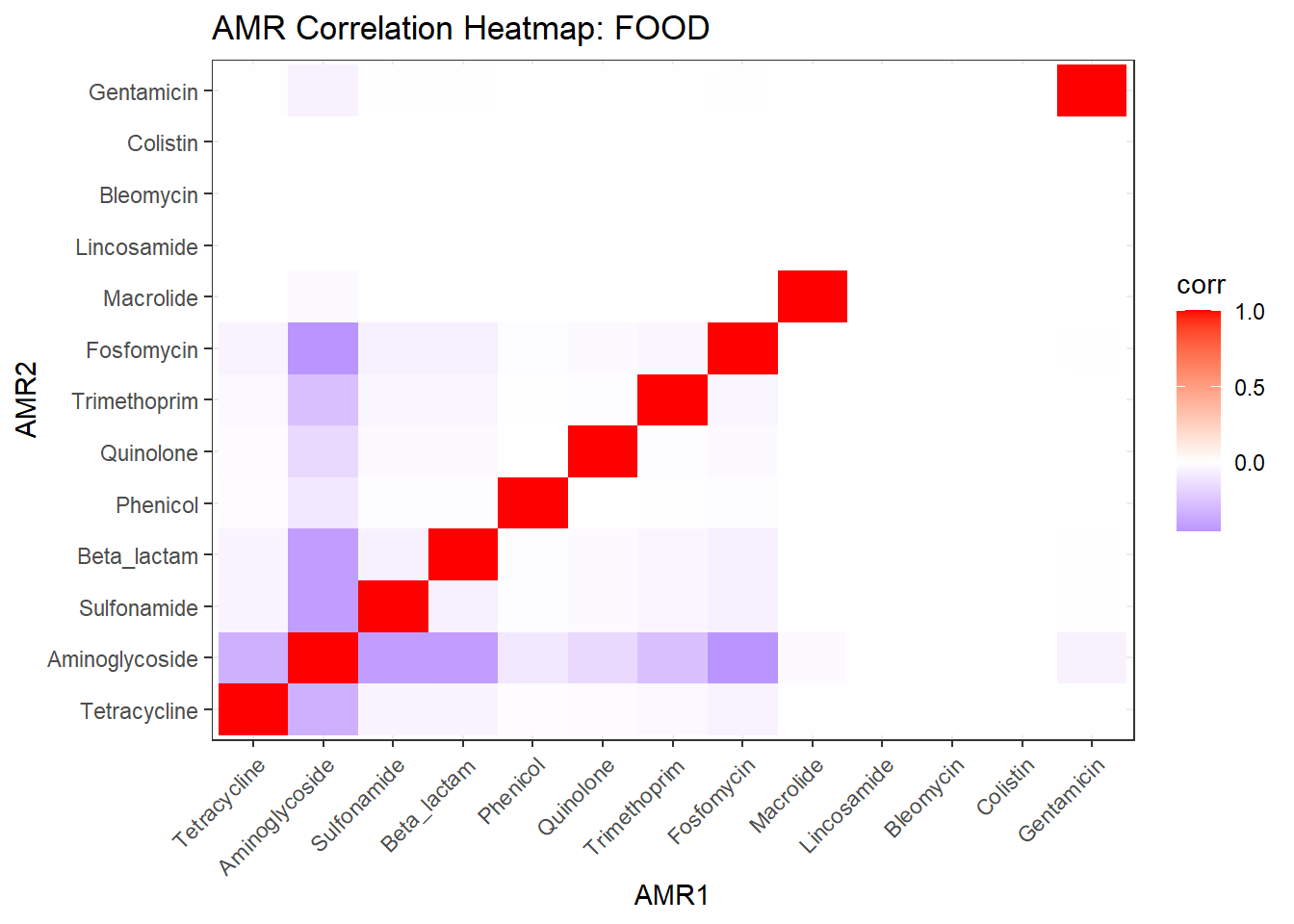
# Publication-ready Heatmap
ggplot(cor_df, aes(AMR1, AMR2, fill = corr)) +
geom_tile(color = "white") +
scale_fill_gradient2(low = "#2166ac", mid = "#f7f7f7", high = "#b2182b",
midpoint = 0, limit = c(-1, 1), name = "Pearson\nCorr") +
theme_minimal() +
theme(axis.text.x = element_text(angle = 45, vjust = 1, hjust = 1),
panel.grid.major = element_blank()) +
labs(title = "Co-occurrence of Antimicrobial Resistance Classes",
x = NULL, y = NULL) -> amr_heatmap_pub
ggsave(here("results/figures/amr_correlation_heatmap.png"), plot = amr_heatmap_pub, dpi = 300, width = 9, height = 7)